In the early days of the pandemic, I wrote squashinformr, an R package for web scraping professional squash data. I’m quite proud of the package, as it was my first real effort in creating open source. But when I wrote the initial version, I committed many cardinal programming sins.
At the time, my approach to writing code was very ‘script-like’. I kept code for my analyses in single Rmarkdown files that I would run from top to bottom. It’s clear looking at old squashinformr code that I expected functions to behave the same way. That is, I wrote the functions as if they were individual scripts that had no common behaviour or tasks. In reality, some functions contained identical code except for a few substituted variables. In short, the code was a mess and vulnerable to bugs.
Since then, I’ve learned software development best practices with the help of a mentor at work. Feeling more confident about my ability to write stable code, I refactored squashinformr earlier this year. I’d like to break down examples of where I applied these best practices as if I were speaking to my past self. If you haven’t heard of these rules, I recommend you apply some of them in your work. That way, you or someone else does not have to rewrite your code in the future 😄
Determine requirements by drawing a diagram
It’s easy to get inspired about a project idea and want to dive straight in to making it. This happened to me when I wrote squashinformr. I was excited by the idea of writing an R package and I had already written code that achieved some of the basic functionality. So I just started writing code and putting hours of time into something I had not thought through. To quote Jurassic Park:
Yeah, yeah, but your scientists were so preoccupied with whether or not they could that they didn’t stop to think if they should.
In other words, I started writing code before determining what the end product should be. I didn’t think about how many functions I would be writing or what they should deliver. I didn’t consider how those functions could share subtasks so I didn’t write helper functions. I ended up wasting a lot of time because of this mistake. To be honest, if I had determined the package’s requirements beforehand, I wouldn’t have written this blog post.
Instead, I could have achieved a high-level vision of the package by drawing a diagram. Using three squashinformr functions as an example, the diagram could look like this:
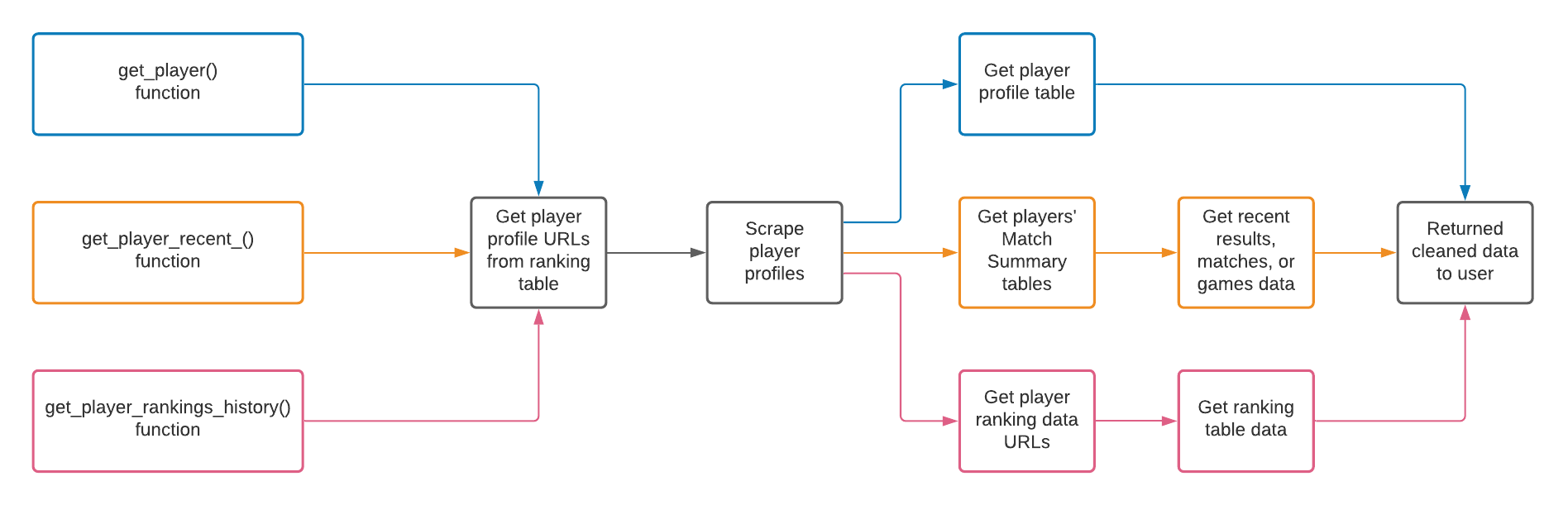
By creating this diagram, I’ve forced myself to think about how the final product works. I’m forced to think about the package functions as a whole and the steps that each function takes towards its desired behaviour. I also have to consider which functions share logic and which logic is unique (e.g. data cleaning). This exercise is informative even if you are the one who came up with the project idea. It helps you determine the scope of the project and leads you toward a refined vision of what you’re about to do.
Development teams will have their own processes for determining requirements, which may be too formal for some personal projects. That’s why I recommend this method. You can decide how formal or detailed you want to be so you don’t exhaust your inspiration in the planning stage. At the same time, you don’t dive in and hope for the best. At the very least, you have an image you can reference to keep you on track with the high-level purpose of your project.
Beware copy + paste
This is a tip that you might have heard before and, like me, ignored out of convenience. Copying and pasting code is a quick and easy solution in the moment. Unfortunately, it is also a sure way to accrue technical debt. It is an easy way to implement the logic you need, but it comes with a catch: it (at least) doubles the length of your code. I made this mistake when writing squashinformr many times over by doing something like this:
get_some_data <- function(category = c("mens", "womens")) {
if (category == "mens") {
## [Code to get men's data]
## [Data cleaning]
}
else if (category == "womens") {
## [Code to get women's data]
## [Data cleaning]
}
## [Common data operations]
return(result)
}
Andrew Tannenbaum explained the consequence of lengthening your code:
In addition, a substantial number of the problems caused by buggy software, which occurs because vendors keep adding more and more features to their programs, which inevitably means more code and thus more bugs.
My pseudo-code function above is waiting for a bug to ruin it. In this scenario, the code that retrieves the men’s data is nearly identical to the code that retrieves the women’s data. If I want to introduce a change or if a bug stops the function from working, I would have to rewrite code in up to four separate places to ensure consistent results. It’s a similar story when it comes to code commenting. I would have to rewrite the comments in up to four places if they were common between each workflow. This is why it’s in our best interest to avoid copying and pasting code. Not only does it make code more vulnerable, but it also creates more work for you in the future.
So what is the alternative strategy to copying and pasting? Writing functions. When you are about to write code that repeats the same logic as other code you’ve written, it’s time to consider whether you should start writing a function. Hadley Wickham wrote a good rule of thumb for this in his R for Data Science book:
You should consider writing a function whenever you’ve copied and pasted a block of code more than twice (i.e. you now have three copies of the same code).
While refactoring squashinformr, I wrote new helper functions that achieved subtasks within the package’s main functions. The result looks something like this:
get_some_data <- function(category = c("mens", "womens")) {
if (category == "mens") {
result <- helper_function(category = category)
}
else if (category == "womens") {
result <- helper_function(category = category)
}
## [Common data operations]
return(result)
}
helper_function <- function(category = c("mens", "womens", "both")) {
## [Code to get data, given category]
## [Data cleaning]
return(data)
}
The code that gets the data is now abstracted away from the main function. The result is succinct code that achieves the desired result without accruing technical debt.
By avoiding copying and pasting, we can make our code more robust and maintainable for the future.
Continuous testing
One thing I did well in the first version of squasinformr was testing. I wrote two main types of tests for every function using the testthat package. Equality tests (i.e. “this function with these inputs should return exactly this result”) and input error tests (e.g. “this function with these inputs should return an error”). These tests would tell me if a function was behaving as expected or return an error if it wasn’t. So all was good while I developed the package, but I forgot to consider something important. How was I going to continue to test these functions after I released squashinformr to the world?
After releasing a package, you take on responsibility for it. Software packages need maintenance and you are the person people go to if it has issues. So, it’s best that you are the first person to know when something is wrong. This is where continuous testing comes in. It’s great to write tests, but it’s not enough to run them every once in a while. You need to run tests on an interval that make sense. If your software is niche and not integral to other people’s work, you can afford to test less often. If it is important and widely used, you should be testing constantly.
So how do you conduct tests on a regular interval? There are many options for software testing at an entreprise level, but only one stand-out solution for everyday open source contributors. GitHub Actions is a flexible workflow system that can be configured to test almost any piece of software. If you host your project on GitHub, automated testing workflows can be achieved by adding a configuration file to a .github/workflows/ directory in your project. The good news is that the GitHub community already hosts useful preconfigured actions that complete all sorts of tasks “out of the box”. For R users, the r-libs/actions repository serves as an extraordinary resource, particularly for testing. If you’re feeling adventurous, GitHub also provides tools to create your own custom actions.
I implemented a modified version of the Standard CI Workflow into squashinformr. In addition to every time I submit a change to the master branch, GitHub Actions runs my tests on squashinformr every 24 hours. This ensures that I won’t go more than a day without knowing something is wrong with the package. I also run these tests in a “matrix configuration”, which is a fancy way of saying “across many operating systems”. So I know my package is accessible to as many users as possible. Now that I’ve taken these precautions, squashinformr is more maintainable than ever. Barring any wild bus accidents, I’ll be able to identify and fix issues as they arise.
Conclusion
These are the three best practices I’ve been able to apply to my open source work with squashinformr. I determined software requirements by drawing high-level diagrams of the finished product. I reduced code redundancy and complexity by writing helper functions. Lastly, I integrated continuous testing through GitHub Actions to ensure timely maintenance and wider accessibility.
After all that, my work is not finished. squashinformr is not perfect, it still needs help! If you are looking to get involved in open source, check out the GitHub repository if this project interests you. Feel free to open an issue or send me an email (hmd@needleinthehay.ca) with your idea. Thanks for reading!